1.
Introduction
Some proteins are considered cytotoxic because their overexpression causes cellular growth defects. However, toxicity is a relative quantity that can only be defined by comparison. The expression limit of a protein, namely, the protein expression level that results in growth defects, may be considered as an index for protein cytotoxicity [1]. In the model eukaryote Saccharomyces cerevisiae, fluorescent proteins and glycolytic enzymes have the highest expression limits, which means that they are the least cytotoxic in this organism [2], while reports suggest that fluorescent proteins could be cytotoxic in other cells [3],[4]. The cytotoxicity of fluorescent proteins can be increased (i.e., their expression limits are decreased) by adding localization signals or mutations that cause misfolding [5],[6]. The mechanism determining the expression limit of a non-toxic protein is recognized as the “protein burden/cost effect,” in which cellular protein production resources are overloaded by extreme overexpression of the protein [1],[7]. In S. cerevisiae, the limit that triggers protein burden is about 15% of the total protein content [2].
The nucleotide sequence that corresponds to the N-terminal of the protein strongly affects the expression level of the protein by modulating the efficiency of translation [8]. The N-terminal amino acid sequences of proteins also affect the stability and thus the expression levels of the protein [9]. N-terminal amino acid sequences also affect the cytotoxicity of proteins because they often transmit localization signals [5]. Although there are several systematic analyses revealing the relationship between amino acid sequences and expression levels [9],[10], predicting the effect of the N-terminal sequence on the expression level and cytotoxicity of the protein remains challenging.
In this study, the effects of the N-terminal sequence on the expression level and cytotoxicity of green fluorescent protein (GFP) were systematically analyzed. Random 30 nucleotides (i.e., ten amino acids) were inserted into the coding sequence for the N-terminal region of GFP. This length was chosen because peptide sequences shorter than ten amino acid length could be linkers connecting different proteins [11], protein localization signals [12]–[14], and protein tags [15],[16] used for protein engineering. GFP consists of a beta-barrel structure as its main tertiary structure [17], and additional N-terminal sequences should be located outside of the main structure.
The effects of the sequence on the expression level and GFP cytotoxicity were assessed using the expression per gene (EPG) and gene copy number limit (CNL) relationship. Several sequences that would decrease the expression level or increase the cytotoxicity were isolated. The lower expression level was associated with codon optimality, and cytotoxicity was increased by the existence of hydrophobic or cysteine residues.
2.
Materials and Method
2.1. Strain, plasmid, media, and growth conditions
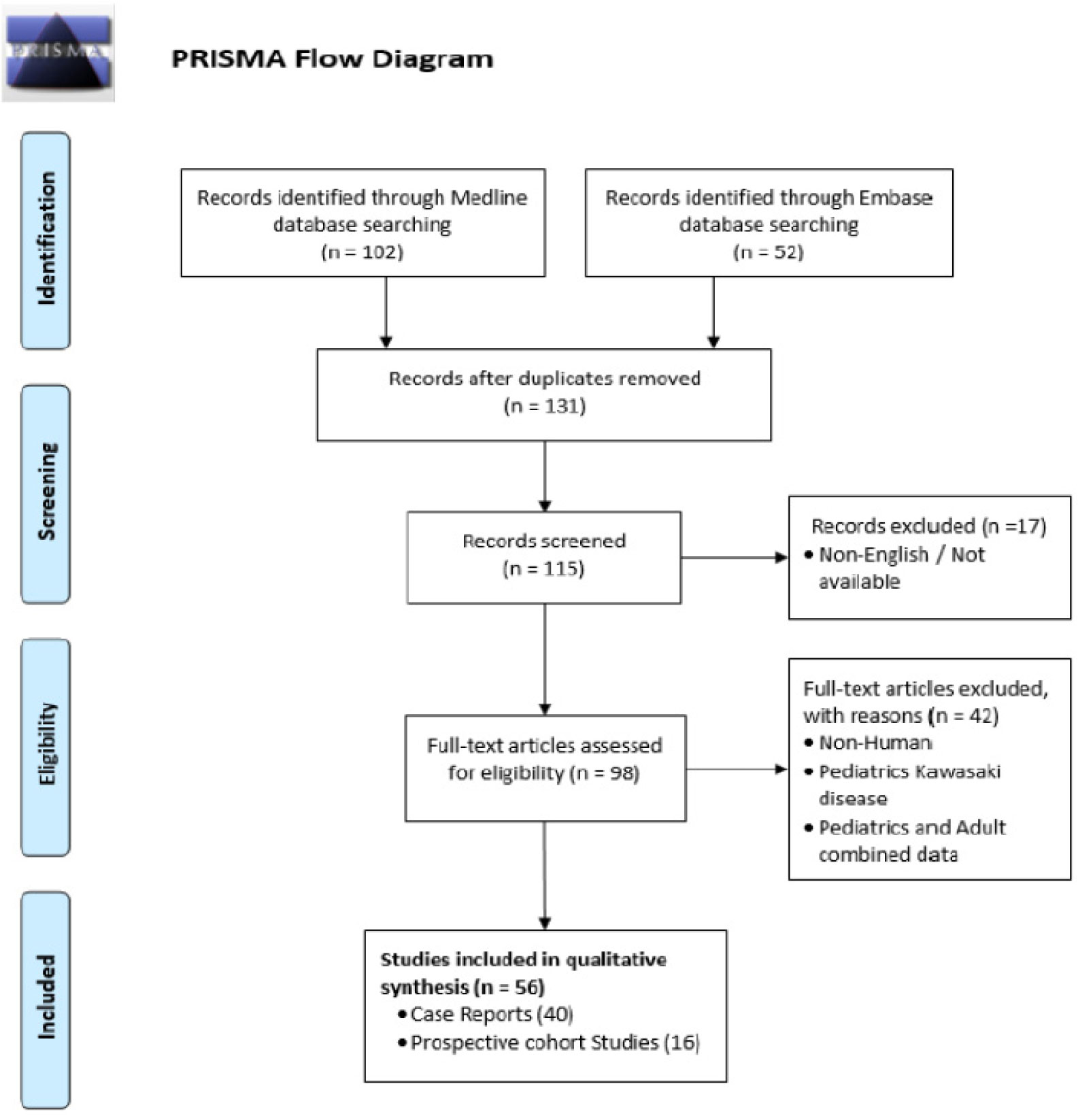
The S. cerevisiae strain BY4741 (MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0) [18] was used in all experiments. pTOW40836-TDH3pro-yEGFP (Figure 1A) [19] was used as the base plasmid. The cultivation and transformation of yeast were performed as previously described [20]. Saccharomyces cerevisiae cells were grown in synthetic complete (SC) medium without uracil (−Ura) or without leucine and uracil (−Leu/Ura) conditions. The SC medium was made using the yeast nitrogen base (4027–512, MP Biomedicals), glucose (16806–25, Nacalai tesque), the –His/–Leu/–Ura drop-out mix (630423, Clontech), histidine (18116–92, Nacalai tesque), and leucine (20327–75, Nacalai tesque).
2.2. Preparation of yeast strains with N10-GFP plasmids
Two DNA fragments were amplified with the primers OHM1002 (5′-CATTTTGTTTGTTTATGTGT-3′)–OSBI851 (5′-ATGAGTATTCAACATTTCCG-3′) and OHM600 (5′-ATGTCTAAAGGTGAAGAATT-3′)–OSBI850 (5′-TTACCAATGCTTAATCAGTG-3′) using pTOW40836-TDH3pro-yEGFP as a template. The N10 DNA fragment was prepared via single-cycle amplification of the template DNA (OHM1000: 5′-TAAACACACATAAACAAACAAAATGNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNTCTAAAGGTGAAGAATTATTCACTG-3′) using the primer OHM1001 (5′-CAGTGAATAATTCTTCACCTTTAGA-3′). The DNA fragments were amplified using KOD-Plus-NEO (KOD-401, TOYOBO). All three DNA fragments were simultaneously introduced into BY4741 cells to join them, and the transformants were cultured on an SC–Ura plate. The N10-GFP plasmids were constructed by homologous recombination-based cloning in vivo [21]. Randomly-chosen 94 colonies of the transformants were picked-up and used for further analysis.
2.3. Measurement of growth rate, GFP fluorescence, and plasmid copy number
Each yeast strain was cultivated in 200 µL SC–Ura media in each well of a 96-well plate (161093, Thermo) at 30 °C for overnight to prepare the preculture. The precultured cells (5 µL each) were transferred into 200 µL of SC–Ura or SC–Leu/Ura medium in 96-well plates, and grown at 30 °C without agitation. The cell density (absorbance 595 nm) and GFP fluorescence (485 nm excitation – 535 emission) were measured every 30 min for 50 hours using the Infinite F200 microplate reader (TECAN). The maximum growth rate (min −1) during the 270 min interval was calculated from the cell density measurements, and the maximum GFP fluorescence (AU) was obtained from the GFP fluorescence measurements. After cell cultivation, cells were collected, and the plasmid copy number in the cell (copies per cell) was measured via real-time PCR as described previously [22]. The expression per gene (EPG, AU) of each N10-GFP was calculated by dividing the maximum GFP fluorescence in SC–Ura by the plasmid copy number in SC–Ura. The plasmid copy number in SC–Leu/Ura was considered as the copy number limit (CNL) of each N10-GFP as the reason described in Results.
2.4. Bioinformatic analysis
The folding energy of mRNA ΔG (kcal/mol) containing the N10 sequence (from −4 nt to +37 nt) was calculated using the mfold web server (http://unafold.rna.albany.edu/?q=mfold). Codon optimality was calculated as the tRNA adaptation index (tAI) using the stALcalc (http://tau-tai.azurewebsites.net/). The existence of a localization signal was predicted using DeepLoc-1.0 (http://www.cbs.dtu.dk/services/DeepLoc/). The GRAVY value was calculated using the GRAVY calculator (http://www.gravy-calculator.de/). The distribution of expected GRAVY values in the 10 amino acid sequences translated from random 30 nucleotide sequences (mean = −0.254 and standard deviation = 0.950) were estimated from the 100,000 randomly generated sequences. The existence of the transmembrane domain (TM) and secondary structure in the N-terminal 30 a.a. were predicted using TMHMM server v. 2.0 (http://www.cbs.dtu.dk/services/TMHMM/) and Jpred4 (http://www.compbio.dundee.ac.uk/jpred/), respectively. The statistical analyses were performed using Microsoft Excel (version 16.36).
3.
Results
3.1. Isolation of N-terminal sequences that modulate the expression level and GFP cytotoxicity
Figure 1A shows the plasmid used to assess the effects of the N-terminal sequences on the expression level and GFP cytotoxicity. A yeast codon-optimized enhanced GFP (yEGFP3) [23] was the GFP used. After the N-terminal methionine, random 30 nucleotides encoding 10 amino acids (N10 sequences) were inserted by homologous recombination-based cloning in S. cerevisiae. The resulting N10-GFPs were expressed under the control of the strong TDH3 promoter (TDH3pro). pTOW40836 was used as a backbone plasmid. The copy number of pTOW40836 in S. cerevisiae cells cultured without leucine and uracil (−Leu/Ura) reflects the CNL of the gene. Under such conditions, the overexpressed gene product inhibits the growth of the cell owing to the genetic tug-of-war (gTOW) effect [22],[24].
The principle of how CNL is determined by the gTOW is explained as follows (Figure 1B). A target gene is cloned into a plasmid for gTOW (Step 1 in Figure 1B), and is used to transform an ura3Δ leu2Δ strain (Step 2 in Figure 1B). In this study, we used TDH3pro_N10-GFP, pTOW40836, and BY4741 as the target, the plasmid for gTOW, and the host strain, respectively. Because of 2µ ORI on the plasmid, the copy number of the plasmid becomes multicopy and diverse among the population. When the cells are transferred into the –Leu/Ura media, the plasmid copy number becomes >100 copies per cell (Step 3 in Figure 1B). This is performed due to leu2-89 on the plasmid, which is an allele of LEU2 harboring a large deletion within its promoter and thus weakly expressed. A higher copy number of leu2-89 is required to fulfill the demand of Leu2 protein to synthesize an essential nutrient leucine (red line in Figure 1B). However, if the target gene has the expression limit to cause cellular defects, the gene copy number and thus the plasmid copy number should be less than the limit, because overexpression of the target beyond the limit causes growth defects (blue line in Figure 1B). The resulting tug-of-war of selection biases between leu2-89 and the target gene (blue and red arrows in Figure 1B) determines the plasmid's copy number, which is determined by the real-time PCR (Step4 in Figure 1B). Because the bias of leu2-89 is always the same, the copy number should reflect the copy number limit (CNL) of the target gene.
Our framework for identifying N10 sequences affecting the expression level and GFP cytotoxicity is explained as follows. The GFP EPG and CNL of the gene should be inversely proportional because the expression limit of GFP (EPG × CNL) is constant; for example, if there are three genes encoding the same GFP, but their EPGs are low, medium, and high, their CNLs should be high, medium, and low, respectively (Figure 2A). The inversely proportional curve is considered as the frontier line in the EGP–CNL space made by the expression limit of GFP (Figure 2B). If an N10 sequence simply changes the EPG of the N10-GFPs, but not its cytotoxicity and expression limit, the EPG–CNL relationship should be on the same frontier line. Conversely, if an N10 sequence increases the cytotoxicity, thus reducing the expression limit of N10-GFPs, the EPG–CNL relationship should shift to the left (Figure 2B, white arrow). Therefore, N10 sequences, which may alter the expression level and GFP cytotoxicity, can be systematically isolated by analyzing the EPG–CNL relationship.
Overall, 94 yeast colonies containing the N10-GFP plasmid were randomly isolated. Then, their growth rates, level of fluorescence, and plasmid copy numbers without uracil (−Ura) and −Leu/Ura conditions (Table S1) were measured. If analyzed N10-GFPs are under the gTOW regime described above, the growth rates and copy numbers in –Leu/Ura conditions should be on the linear red line created by the relationship between the growth rate and copy number of leu2-89 (Figure 1B). The relationship between the growth rates and copy numbers among the 94 isolates was r = 0.82 (Table S1), suggesting that they are under the gTOW regime, and thus the plasmid copy number under −Leu/Ura conditions was considered as the gene CNL. The GFP EPG was calculated by dividing the level of GFP fluorescence by the plasmid copy number. To calculate the EPG, the level of GFP fluorescence and plasmid copy number obtained under −Ura conditions were used, wherein GFP overexpression produces less growth defects. As shown in Figure 3A, the 94 isolates exhibited various EPG–CNL relationships, which indicates that the N10 sequences modulated the expression levels and toxicities of GFP. Importantly, the EPG–CNL relationships created a frontier line (dotted line in Figure 3A) as it is theoretically predicted in Figure 2B. Most of the N10-GFPs were located around the frontier line, suggesting that most N10 sequences just modulate the expression level of GFP but not the cytotoxicity. Sixteen representative isolates were selected (Figure 3A), and the N10 sequences of each were determined. Seven isolates (#6, #44, #2, #20, #90, #1, and #85) were on the frontier line, but their locations were different, which suggests that their N10 sequences modulated the expression levels of GFP. Four isolates (#15, #75, #80, and #82) were located inside the frontier line, which suggests that their N10 sequences were cytotoxic. Three toxic non-fluorescent isolates (#51, #52, and #64) and two non-toxic non-fluorescent isolates (#55 and #74) were also analyzed.
Table 1 presents the sequencing results. First, toxic non-fluorescent isolates (#51, #52, and #64), but not N10-GFPs, contained the TDH3 gene. Therefore, the plasmids were probably constructed through homologous recombination between the base plasmid and TDH3 within the S. cerevisiae genome. Because Tdh3 has a lower expression limit than GFP [19], isolates expressing Tdh3 should have exhibited EPG–CNL relationships that differed from those of GFP. Second, non-toxic non-fluorescent isolates (#55 and #74) had nonsense codons within their N10 sequences, indicating that the genes do not encode any proteins. Third, isolates showing the highest EPGs (#6 and #44) did not contain the N10 sequence but rather GFP alone; these isolates were probably constructed by self-ligation of the base plasmid. Because #6 and #44 are located on the top of the frontier line, inserting an N10 sequence seemed to reduce the expression level or increase the GFP cytotoxicity. The other nine isolates contained various N10 sequences; among these, five (#2, #20, #90, #1, and #85) decreased the expression level, whereas four (#15, #75, #70, and #82) increased cytotoxicity as shown in Figure 3A. Figure 3B shows the CNLs and EPGs of the representative isolates.
3.2. Codon optimality of the N10 sequence affects the expression level
To reveal the sequence property that determines the expression level of N10-GFPs, the mRNA folding energy [25] and codon optimality (tRNA adaptation index; tAI) [26] were calculated (Table S2). The existence of localization signals [27] was also predicted. Although no significantly associated values with the expression level and cytotoxicity were noted, the tAIs of isolated N10 sequences were found to be lower than those of GFP (0.320). Their codon usage was thus optimized to that of the highly expressed TDH3. Codon optimization altered the EPG–CNL relationship profiles of N10-GFPs (Figure 4). Most importantly, the optimization increased the EPGs of isolates with low expression levels of N10-GFPs (#1 and #85), which suggests that codon optimality is responsible for their low EPGs. Thus, codon optimality of the N10 sequences seemed to be the most important contributing factor to the expression levels of N10-GFPs.
3.3. Existence of hydrophobic and cysteine residues in the N10 sequence might increase N10-GFP cytotoxicity
Next, the relationship between the amino acid composition in the N10 sequence and N10-GFP cytotoxicity was analyzed. First, the grand average of hydropathy (GRAVY) value of the amino acid sequence [28] (Table S2) was calculated, which has a positive relationship with hydrophobicity. The toxic N10 sequences had significantly higher GRAVY values than those of the non-toxic N10 sequences (p = 0.028, Student t-test, Figure 5A). The probability of obtaining four N10 sequences with GRAVY values of >0.696 is <0.0006, which suggests that hydrophobic residues contribute to the cytotoxicity of the N10 sequences. Next, the frequency of each amino acid in the four toxic N10 sequences (#15, #75, #80, and #82) was calculated and compared with the expected frequencies in the codon table (Figure 5B). The quantities of cysteine, methionine, and tryptophan residues were over two-fold of the expected frequencies. The p-value of the appearance of cysteine was 0.006 according to the results of the chi-square test, which was slightly insignificant under the false discovery rate <0.05 (p < 0.003). Therefore, we concluded that cysteine residues were over-represented in toxic N10-GFPs. Structural analyses of the N-terminal 30 a.a. containing N10 sequences and the N-terminal of GFP were also performed (Table S2). A transmembrane domain prediction by TMHMM (v2.0) [29] did not give any transmembrane domain in the sequences, and a secondary structure prediction by Jpred4 [30] gave some structures, but there was no structural consistency within the toxic and non-toxic N10 sequences.
4.
Discussions
The nucleotide sequence encoding the N-terminal of a protein plays a substantial role in protein production because it is where ribosomes are assembled and translation is initiated. The secondary structure of mRNA and codon optimality affect the initiation of translation [8]. In this study, it was confirmed that codon optimality was a major factor in reducing the expression level of GFP, although mRNA stability did not affect the expression level. A question that was raised during this study focused on the mechanism causing cytotoxicity of N10 sequences. A possible explanation for this phenomenon is the artificial creation of localization signals; this occurs because the addition of localization signals at the N-terminal of GFP increases its cytotoxicity [5]. However, three out of the four toxic N10 sequences did not have any signature localization signal. Moreover, no localization of toxic N10-GFPs (data not shown) was observed. Agammaegative sequences also increase the cytotoxicity of proteins [31]; however, no agammaegation of toxic N10-GFPs (data not shown) was noted. Although a small subset of toxic N10 sequences was analyzed here, they showed statistically significant concentration of hydrophobic residues and a preferential appearance of cysteine residues. Overexpression of cysteine-containing glycolytic enzymes constitutes the agammaegates connected by the S–S bond [2],[32], and removal of cysteine from Tpi1 increases its expression limit [2]. Therefore, the existence of unimportant cysteines might increase the cytotoxicity of highly expressed proteins regardless of their positions. Hydrophobic residues seemed to increase the GFP cytotoxicity. This may be owing to non-specific hydrophobic interactions, and therefore, they might not be preferred in the outside of the main tertiary structure of proteins. Another possible mechanism to cause cytotoxicity is the perturbation of translation. Sequences rich in proline and positively-charged residues cause slowdown in translational elongation [33]. However, this might not be the case in this analysis because any enrichment of proline or positively-charged residues (lysine or arginine) was observed in the toxic N10 sequences (Figure 5B).








 DownLoad:
DownLoad: