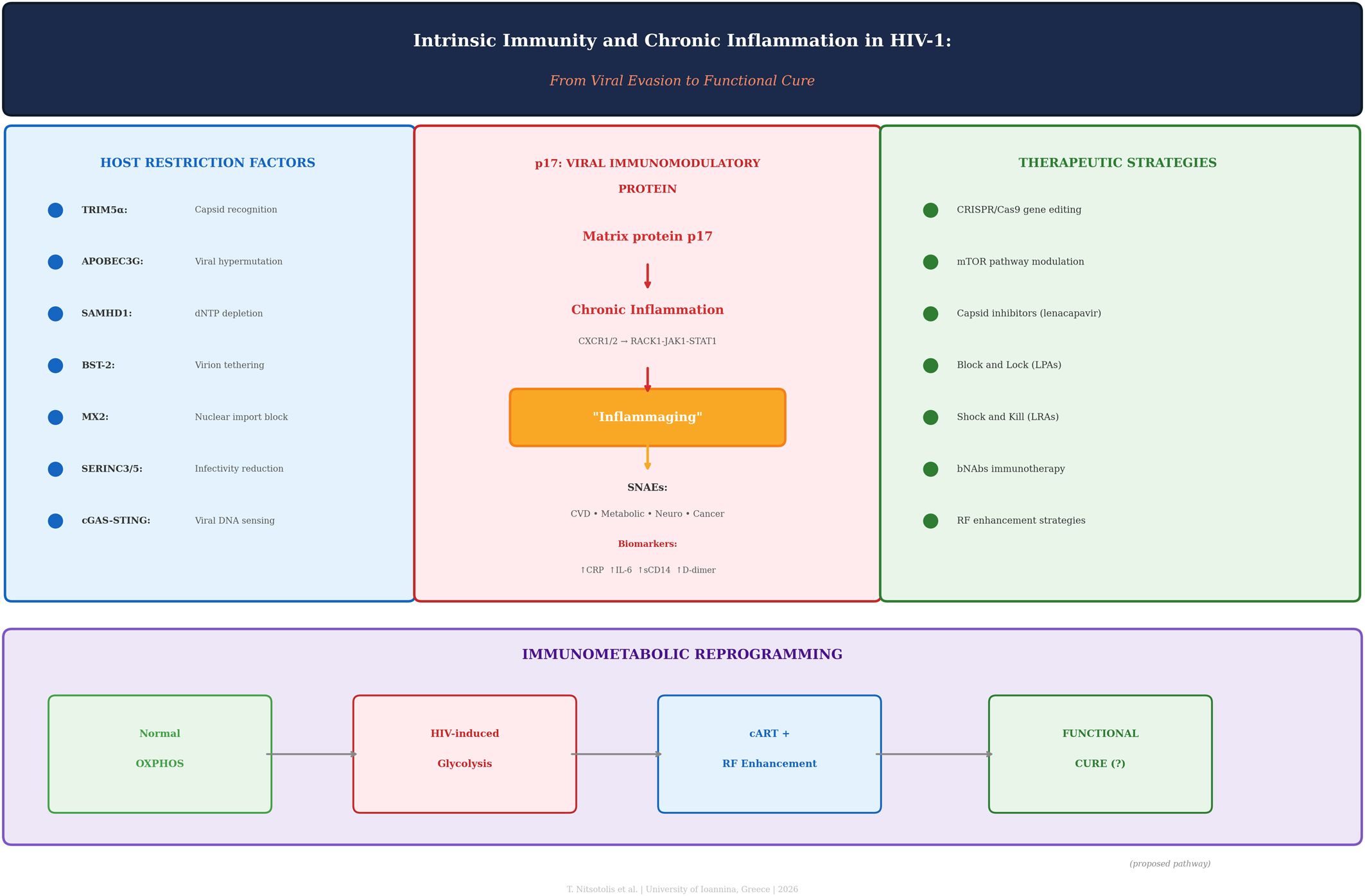
Despite the widespread success of combination antiretroviral therapy (cART) in suppressing plasma viremia to undetectable levels, people living with HIV-1 (PLWH) continue to face a significantly elevated risk of chronic inflammation and Serious Non-AIDS Events (SNAEs). In this narrative review, we bridge the critical gap between molecular virology, immunometabolism, and clinical pathology by examining the complex interface of intrinsic immunity and viral persistence. We analyzed the evolutionary “arms race” between conserved host restriction factors, including TRIM5α, APOBEC3G, SAMHD1, BST-2, MX2, and SERINC, and the sophisticated viral evasion mechanisms that facilitate reservoir establishment. We further examined the role of bacterial translocation and gut barrier dysfunction in perpetuating systemic inflammation, emphasizing how HIV-1-mediated depletion of mucosal Th17 cells and disruption of tight junction proteins create a “leaky gut” that permits microbial product translocation despite suppressive therapy. Among viral proteins that may contribute to residual pathology during suppressive cART, we focused on the HIV-1 matrix protein p17, which has been proposed to function as a secreted “viral cytokine” from latent reservoirs, acting through CXCR1/CXCR2 receptors and the RACK1-JAK1-STAT1 pathway. Although primarily characterized in in vitro and ex vivo models, emerging data suggested that p17 may sustain systemic immune activation and metabolic reprogramming; however, its relative contribution compared with other viral proteins (Tat, Nef, gp120) in virologically suppressed patients remains to be fully delineated in human studies.
Furthermore, we examined how HIV-1 hijacks cellular bioenergetics by shifting host cells from oxidative phosphorylation to aerobic glycolysis. We present an integrative model that connects restriction factor biology, p17-mediated chronic inflammation, immunometabolic dysregulation, and gut barrier dysfunction into a unified pathogenic framework, distinguishing established mechanisms from working hypotheses. Last, we assessed emerging therapeutic strategies, including CRISPR/Cas9-mediated enhancement of restriction factors, modulation of the mTOR pathway, and novel “Shock and Kill” approaches, stratified by development stage and demonstrated endpoints, offering potential pathways toward a functional cure.
Citation: Thomas Nitsotolis, Stelios F Assimakopoulos, Elli Kouriannidi, Maria Lagadinou, Alexia Papalexandrou, Petros Ioannou, Markos Marangos, Haralampos Milionis, Eirini Christaki. Host restriction factors and p17-Driven inflammaging in HIV-1: From molecular pathogenesis to functional cure[J]. AIMS Microbiology, 2026, 12(1): 92-125. doi: 10.3934/microbiol.2026004
Despite the widespread success of combination antiretroviral therapy (cART) in suppressing plasma viremia to undetectable levels, people living with HIV-1 (PLWH) continue to face a significantly elevated risk of chronic inflammation and Serious Non-AIDS Events (SNAEs). In this narrative review, we bridge the critical gap between molecular virology, immunometabolism, and clinical pathology by examining the complex interface of intrinsic immunity and viral persistence. We analyzed the evolutionary “arms race” between conserved host restriction factors, including TRIM5α, APOBEC3G, SAMHD1, BST-2, MX2, and SERINC, and the sophisticated viral evasion mechanisms that facilitate reservoir establishment. We further examined the role of bacterial translocation and gut barrier dysfunction in perpetuating systemic inflammation, emphasizing how HIV-1-mediated depletion of mucosal Th17 cells and disruption of tight junction proteins create a “leaky gut” that permits microbial product translocation despite suppressive therapy. Among viral proteins that may contribute to residual pathology during suppressive cART, we focused on the HIV-1 matrix protein p17, which has been proposed to function as a secreted “viral cytokine” from latent reservoirs, acting through CXCR1/CXCR2 receptors and the RACK1-JAK1-STAT1 pathway. Although primarily characterized in in vitro and ex vivo models, emerging data suggested that p17 may sustain systemic immune activation and metabolic reprogramming; however, its relative contribution compared with other viral proteins (Tat, Nef, gp120) in virologically suppressed patients remains to be fully delineated in human studies.
Furthermore, we examined how HIV-1 hijacks cellular bioenergetics by shifting host cells from oxidative phosphorylation to aerobic glycolysis. We present an integrative model that connects restriction factor biology, p17-mediated chronic inflammation, immunometabolic dysregulation, and gut barrier dysfunction into a unified pathogenic framework, distinguishing established mechanisms from working hypotheses. Last, we assessed emerging therapeutic strategies, including CRISPR/Cas9-mediated enhancement of restriction factors, modulation of the mTOR pathway, and novel “Shock and Kill” approaches, stratified by development stage and demonstrated endpoints, offering potential pathways toward a functional cure.

adeno-associated virus
antibody-dependent cellular cytotoxicity
antibody-dependent cellular phagocytosis
acquired immunodeficiency syndrome
protein kinase B
adaptor protein 2
apolipoprotein B mRNA editing enzyme catalytic polypeptide-like 3G
analytical treatment interruption
blood-brain barrier
broadly neutralizing antibody
bone marrow stromal antigen 2
capsid
combination antiretroviral therapy
CRISPR-associated protein 9 endonuclease
C-C chemokine receptor type 5
cluster of differentiation
cyclin-dependent kinases 1 and 2
cyclic GMP-AMP
cyclic GMP-AMP synthase
chronic kidney disease
central nervous system
cleavage and polyadenylation specificity factor 6
circulating recombinant form
clustered regularly interspaced short palindromic repeats
C-terminal domain
cytotoxic T lymphocyte
C-X-C motif chemokine receptor 1
C-X-C motif chemokine receptor 2
C-X-C chemokine receptor type 4
cyclophilin A
DDB1 and CUL4 associated factor 1
deoxyribonucleoside triphosphate
extracellular matrix
envelope glycoprotein
endoplasmic reticulum
farnesoid X receptor
group-specific antigen
gut-associated lymphoid tissue
guanosine monophosphate
glycoprotein 41
glycoprotein 120
glycoprotein 160
glycosylphosphatidylinositol
guide RNA
HIV-associated neurocognitive disorders
hepatocellular carcinoma
histone deacetylase inhibitor
human immunodeficiency virus
heptad repeat 1
heptad repeat 2
hematopoietic stem and progenitor cell
heparan sulfate proteoglycans
intestinal fatty acid-binding protein
intercellular adhesion molecule 1
interferon gamma inducible protein 16
interferon
interleukin-17
interleukin-22
integrase
interferon regulatory factor 3
immune reconstitution inflammatory syndrome
interferon-stimulated gene
Janus kinase 1
junctional adhesion molecules
lipopolysaccharide-binding protein
lipid nanoparticle
latency-promoting agent
lipopolysaccharide
latency-reversing agent
long terminal repeat
matrix protein
mitogen-activated protein kinase
metabolic dysfunction-associated steatotic liver disease
monocyte chemoattractant protein 1
major histocompatibility complex
myosin light chain kinase
matrix metalloproteinase 2/9
mechanistic target of rapamycin
mTOR complex 1
myxovirus resistance protein 2
nucleocapsid
negative regulatory factor
nuclear factor kappa B
natural killer
nuclear pore complex
N-terminal domain
open reading frame
oxidative phosphorylation
pathogen-associated molecular patterns
plasmacytoid dendritic cell
pre-integration complex
protein kinase C
people living with HIV
polymerase
protein-protein interaction
protease
peroxiredoxin 2
pre-exposure prophylaxis
pattern recognition receptor
receptor for activated C kinase 1
regulator of expression of virion proteins
retinoic acid-inducible gene I
reactive oxygen species
Rev response element
reverse transcriptase
SAM and HD domain-containing protein 1
soluble CD14
short-chain fatty acids
serine incorporator
simian immunodeficiency virus
serious non-AIDS events
signal transducer and activator of transcription 1
stimulator of interferon genes
surface subunit
transactivation response element
transactivator of transcription
T helper 17 cells
Toll-like receptor
transmembrane
tissue macrophage reservoir
tumor necrosis factor alpha
viral infectivity factor
viral protein R
viral protein U
viral protein X
zinc finger antiviral protein
zonula occludens-1
zonula occludens-2
zonula occludens-3
bromodomain-containing protein 4.

| [1] |
Tusch E, Ryom L, Pelchen-Matthews A, et al. (2024) Trends in mortality in people with HIV from 1999 through 2020: A multicohort collaboration. Clin Infect Dis 79: 1242-1257. https://doi.org/10.1093/cid/ciae228 
|
| [2] |
Hileman CO, Funderburg NT (2017) Inflammation, immune activation, and antiretroviral therapy in HIV. Curr HIV/AIDS Rep 14: 93-100. https://doi.org/10.1007/s11904-017-0356-x 
|
| [3] |
Nitsotolis T, Kyriakoulis KG, Kollias A, et al. (2024) Comparison of integrase strand transfer inhibitors (INSTIs) and protease-boosted inhibitors (PIs) on the reduction in chronic immune activation in a virally suppressed, mainly male population living with HIV (PLWH). Medicina (Kaunas) 60: 331. https://doi.org/10.3390/medicina60020331 
|
| [4] |
Prakoeswa FRS, Maharani F, Bestari RS, et al. (2025) Aging and HIV: Recent findings in contributing factors. AIDS Res Treat 2025: 8814760. https://doi.org/10.1155/arat/8814760 
|
| [5] |
Kulpa DA, Paiardini M, Silvestri G (2025) Immune-mediated strategies to solving the HIV reservoir problem. Nat Rev Immunol 25: 542-553. https://doi.org/10.1038/s41577-025-01136-7 
|
| [6] |
Aounallah M, Dagenais-Lussier X, El-Far M, et al. (2016) Current topics in HIV pathogenesis, part 2: Inflammation drives a Warburg-like effect on the metabolism of HIV-infected subjects. Cytokine Growth Factor Rev 28: 1-10. https://doi.org/10.1016/j.cytogfr.2016.01.001 
|
| [7] |
Basting CM, Klatt NR (2025) Dissecting the impact of the gut microbiome on HIV Reservoir dynamics. J Infect Dis : jiaf560. https://doi.org/10.1093/infdis/jiaf560 
|
| [8] |
Serrano-Villar S, Rojo D, Martínez-Martínez M, et al. (2024) HIV-associated gut microbial alterations are dependent on host and geographic context. Nat Commun 15: 1055. https://doi.org/10.1038/s41467-023-44566-4 
|
| [9] |
Thompson CG, Gay CL, Kashuba ADM (2023) The human microbiome and gut–liver axis in people living with HIV. Curr HIV/AIDS Rep 20: 170-180. https://doi.org/10.1007/s11904-023-00657-x 
|
| [10] |
Assimakopoulos SF, Papageorgiou I, Charonis A (2011) Enterocytes' tight junctions: From molecules to diseases. World J Gastrointest Pathophysiol 2: 123-137. https://doi.org/10.4291/wjgp.v2.i6.123 
|
| [11] |
Assimakopoulos SF, Dimitropoulou D, Marangos M, et al. (2014) Intestinal barrier dysfunction in HIV infection: pathophysiology, clinical implications, and potential therapies. Infection 42: 951-959. https://doi.org/10.1007/s15010-014-0666-5 
|
| [12] |
Isaguliants M, Bayurova E, Avdoshina D, et al. (2021) Oncogenic effects of HIV-1 proteins: Mechanisms behind. Cancers (Basel) 13: 305. https://doi.org/10.3390/cancers13020305 
|
| [13] |
Liu Y, Zhu J, Wu W, et al. (2025) HIV-1 matrix protein p17: a key factor in HIV-associated cancers. Front Virol 5: 1584507. https://doi.org/10.3389/fviro.2025.1584507 
|
| [14] |
Caccuri F, Neves V, Gano L, et al. (2022) The HIV-1 matrix protein p17 does cross the blood-brain barrier. J Virol 96: e0120021. https://doi.org/10.1128/jvi.01200-21 
|
| [15] | Messali S, Zani A, Bonfanti C, et al. (2024) Detection of HIV-1 matrix protein p17 in sera of viremic and aviremic patients. J Virol Methods 323: 114846. https://doi.org/10.1016/j.jviromet.2023.114846 |
| [16] |
Renga B, Francisci D, D'Amore C, et al. (2012) The HIV matrix protein p17 subverts nuclear receptors expression and induces a STAT1-dependent proinflammatory phenotype in monocytes. PLoS One 7: e35924. https://doi.org/10.1371/journal.pone.0035924 
|
| [17] |
Peeters M, Jung M, Ayouba A (2013) The origin and molecular epidemiology of HIV. Expert Rev Anti Infect Ther 11: 885-896. https://doi.org/10.1586/14787210.2013.825443 
|
| [18] | Bush S, Tebit DM (2015) HIV-1 group O origin, evolution, pathogenesis, and treatment: unraveling the complexity of an outlier 25 years later. AIDS Rev 17: 147-158. |
| [19] |
Visseaux B, Damond F, Matheron S, et al. (2016) HIV-2 molecular epidemiology. Infect Genet Evol 46: 233-240. https://doi.org/10.1016/j.meegid.2016.08.010 
|
| [20] |
Skittrall JP, Ingemarsdotter CK, Gog JR, et al. (2019) A scale-free analysis of the HIV-1 genome demonstrates multiple conserved regions of structural and functional importance. PLoS Comput Biol 15: e1007345. https://doi.org/10.1371/journal.pcbi.1007345 
|
| [21] |
Nikolaitchik OA, Islam S, Kitzrow JP, et al. (2023) HIV-1 usurps transcription start site heterogeneity of host RNA polymerase II to maximize replication fitness. Proc Natl Acad Sci USA 120: e2305103120. https://doi.org/10.1073/pnas2305103120 
|
| [22] |
Duchon A, Hu WS (2024) HIV-1 RNA genome packaging: it's G-rated. mBio 15: e0086123. https://doi.org/10.1128/mbio.00861-23 
|
| [23] |
Sakuragi J (2011) Morphogenesis of the infectious HIV-1 virion. Front Microbiol 2: 242. https://doi.org/10.3389/fmicb.2011.00242 
|
| [24] |
Parthasarathy D, Pothula KR, Ratnapriya S, et al. (2024) Conformational flexibility of HIV-1 envelope glycoproteins modulates transmitted/founder sensitivity to broadly neutralizing antibodies. Nat Commun 15: 7334. https://doi.org/10.1038/s41467-024-51656-4 
|
| [25] |
Klasse PJ, Sanders RW, Ward AB, et al. (2025) The HIV-1 envelope glycoprotein: structure, function and interactions with neutralizing antibodies. Nat Rev Microbiol 23: 734-752. https://doi.org/10.1038/s41579-025-01206-6 
|
| [26] |
Chiliveri SC, Louis JM, Best RB, et al. (2022) Real-time exchange of the lipid-bound intermediate and post-fusion states of the hiv-1 gp41 ectodomain. J Mol Biol 434: 167683. https://doi.org/10.1016/j.jmb.2022.167683 
|
| [27] |
Chaudhuri E, Jang S, Chakraborty R, et al. (2025) CPSF6 promotes HIV-1 preintegration complex function. J Virol 99: e0049025. https://doi.org/10.1128/jvi.00490-25 
|
| [28] |
Rohlfes N, Radhakrishnan R, Singh PK, et al. (2025) The nuclear localization signal of CPSF6 governs post-nuclear import steps of HIV-1 infection. PLoS Pathog 21: e1012354. https://doi.org/10.1371/journal.ppat.1012354 
|
| [29] |
Zani A, Messali S, Uggeri M, et al. (2024) Detection of HIV-1 matrix protein p17 in sera of viremic and aviremic patients. J Virol Methods 324: 114858. https://doi.org/10.1016/j.jviromet.2023.114858 
|
| [30] |
Bugatti A, Caccuri F, Filippini F, et al. (2021) Binding to PI(4,5)P2 is indispensable for secretion of B-cell clonogenic HIV-1 matrix protein p17 variants. J Biol Chem 297: 100934. https://doi.org/10.1016/j.jbc.2021.100934 
|
| [31] |
Dubé M, Tastet O, Dufour C, et al. (2023) Spontaneous HIV expression during suppressive ART is associated with the magnitude and function of HIV-specific CD4+ and CD8+ T cells. Cell Host Microbe 31: 1507-1522.e5. https://doi.org/10.1016/j.chom.2023.08.006 
|
| [32] |
Caccuri F, Marsico S, Fiorentini S, et al. (2016) HIV-1 matrix protein p17 and its receptors. Curr Drug Targets 17: 23-32. https://doi.org/10.2174/1389450116666150825110840 
|
| [33] |
Caccuri F, Giagulli C, Bugatti A, et al. (2012) HIV-1 matrix protein p17 promotes angiogenesis via chemokine receptors CXCR1 and CXCR2. Proc Natl Acad Sci USA 109: 14580-14585. https://doi.org/10.1073/pnas1206605109 
|
| [34] |
Bugatti A, Paiardi G, Urbinati C, et al. (2019) Heparin and heparan sulfate proteoglycans promote HIV-1 p17 matrix protein oligomerization: computational, biochemical and biological implications. Sci Rep 9: 15768. https://doi.org/10.1038/s41598-019-52201-w 
|
| [35] |
Renga B, Francisci D, D'Amore C, et al. (2012) The HIV matrix protein p17 subverts nuclear receptors expression and induces a STAT1-dependent proinflammatory phenotype in monocytes. PLoS One 7: e35924. https://doi.org/10.1371/journal.pone.0035924 
|
| [36] |
Bugatti A, Marsico S, Mazzuca P, et al. (2020) Role of autophagy in von willebrand factor secretion by endothelial cells and in the in vivo thrombin-antithrombin complex formation promoted by the HIV-1 matrix protein p17. Int J Mol Sci 21: 2022. https://doi.org/10.3390/ijms21062022 
|
| [37] |
Renga B, Francisci D, Carino A, et al. (2015) The HIV matrix protein p17 induces hepatic lipid accumulation via modulation of nuclear receptor transcriptoma. Sci Rep 5: 15403. https://doi.org/10.1038/srep15403 
|
| [38] |
Okine T, Hill E, Sheran K, et al. (2025) Beyond viral suppression: decoding the mitochondrial-immune axis in HIV-associated inflammation and immune dysfunction. Front Cell Infect Microbiol 15: 1686785. https://doi.org/10.3389/fcimb.2025.1686785 
|
| [39] |
Palmer CS, Palchaudhuri R, Albargy H, et al. (2018) Exploiting immune cell metabolic machinery for functional HIV cure and the prevention of inflammaging. F1000Res 7: 125. https://doi.org/10.12688/f1000research.11881.1 
|
| [40] |
Song YH, Kim H, Baur AS, et al. (2025) Uncovering the evolving arms race between host immunity and HIV-1. Trends Immunol 46: 728-740. https://doi.org/10.1016/j.it.2025.08.004 
|
| [41] |
Ganser-Pornillos BK, Pornillos O (2019) Restriction of HIV-1 and other retroviruses by TRIM5. Nat Rev Microbiol 17: 546-556. https://doi.org/10.1038/s41579-019-0225-2 
|
| [42] |
Zhao G, Ke D, Vu T, et al. (2011) Rhesus TRIM5α disrupts the HIV-1 capsid at the inter-hexamer interfaces. PLoS Pathog 7: e1002009. https://doi.org/10.1371/journal.ppat.1002009 
|
| [43] |
Ito F, Alvarez-Cabrera AL, Liu S, et al. (2023) Structural basis for HIV-1 antagonism of host APOBEC3G via Cullin E3 ligase. Sci Adv 9: eade3168. https://doi.org/10.1126/sciadv.ade3168 
|
| [44] |
Iwatani Y, Matsuoka K, Ode H, et al. (2025) The unique structure of the highly conserved PPLP region in HIV-1 Vif is critical for the formation of APOBEC3 recognition interfaces. mBio 16: e0333224. https://doi.org/10.1128/mbio.03332-24 
|
| [45] |
Kouno T, Shibata S, Shigematsu M, et al. (2023) Structural insights into RNA bridging between HIV-1 Vif and antiviral factor APOBEC3G. Nat Commun 14: 4037. https://doi.org/10.1038/s41467-023-39796-5 
|
| [46] |
Zhao Y, Zhao K, Wang S, et al. (2022) Multi-functional BST2/tetherin against HIV-1, other viruses and LINE-1. Front Cell Infect Microbiol 12: 979091. https://doi.org/10.3389/fcimb.2022.979091 
|
| [47] |
McCown C, Yu CH, Ivanov DN (2025) SAMHD1 shapes deoxynucleotide triphosphate homeostasis by interconnecting the depletion and biosynthesis of different dNTPs. Nat Commun 16: 793. https://doi.org/10.1038/s41467-025-56208-y 
|
| [48] |
Larrous P, Garnier C, Morel M, et al. (2025) Deciphering lentiviral Vpr/Vpx determinants required for HUSH and SAMHD1 antagonism highlights the molecular plasticity of these evolutionary conflicts. J Virol 99: e0019825. https://doi.org/10.1128/jvi.00198-25 
|
| [49] |
Flick H, Venbakkam A, Singh PK, et al. (2025) Interplay between the cyclophilin homology domain of RANBP2 and MX2 regulates HIV-1 capsid dependencies on nucleoporins. mBio 16: e0264624. https://doi.org/10.1128/mbio.02646-24 
|
| [50] |
Layish B, Goli R, Flick H, et al. (2024) Virus specificity and nucleoporin requirements for MX2 activity are affected by GTPase function and capsid-CypA interactions. PLoS Pathog 20: e1011830. https://doi.org/10.1371/journal.ppat.1011830 
|
| [51] |
Leonhardt SA, Purdy MD, Grover JR, et al. (2023) Antiviral HIV-1 SERINC restriction factors disrupt virus membrane asymmetry. Nat Commun 14: 4368. https://doi.org/10.1038/s41467-023-39262-2 
|
| [52] |
Yu J, Liu C, Qu X, et al. (2025) SERINC5 counters retroviruses and non-retroviruses. Front Cell Infect Microbiol 14: 1516806. https://doi.org/10.3389/fcimb.2024.1516806 
|
| [53] |
Scott TM, Arnold LM, Powers JA, et al. (2025) Cell-free assays reveal that the HIV-1 capsid protects reverse transcripts from cGAS immune sensing. PLoS Pathog 21: e1012206. https://doi.org/10.1371/journal.ppat.1012206 
|
| [54] |
Hurst TP, Aswad A, Karamitros T, et al. (2019) Interferon-Inducible Protein 16 (IFI16) has a broad-spectrum binding ability against ssDNA targets: An evolutionary hypothesis for antiretroviral checkpoint. Front Microbiol 10: 1426. https://doi.org/10.3389/fmicb.2019.01426 
|
| [55] |
Nchioua R, Kmiec D, Krchlikova V, et al. (2025) Host ZAP activity correlates with the levels of CpG suppression in primate lentiviruses. Proc Natl Acad Sci USA 122: e2419489122. https://doi.org/10.1073/pnas2419489122 
|
| [56] |
Onomoto K, Onoguchi K, Yoneyama M (2021) Regulation of RIG-I-like receptor-mediated signaling: interaction between host and viral factors. Cell Mol Immunol 18: 539-555. https://doi.org/10.1038/s41423-020-00602-7 
|
| [57] |
Wang JW, Liu JH, Xun JJ (2025) CCR5 gene editing and HIV immunotherapy: current understandings, challenges, and future directions. Front Immunol 16: 1590690. https://doi.org/10.3389/fimmu.2025.1590690 
|
| [58] |
Feist WN, Luna SE, Ben-Efraim K, et al. (2025) Multilayered HIV-1 resistance in HSPCs through CCR5 Knockout and B-cell secretion of HIV-inhibiting antibodies. Nat Commun 16: 3103. https://doi.org/10.1038/s41467-025-58371-8 
|
| [59] |
Dudek AM, Feist WN, Sasu EJ, et al. (2024) A simultaneous knockout knockin genome editing strategy in HSPCs potently inhibits CCR5- and CXCR4-tropic HIV-1 infection. Cell Stem Cell 31: 499-518.e6. https://doi.org/10.1016/j.stem.2024.03.002 
|
| [60] |
Khamaikawin W, Saisawang C, Tassaneetrithep B, et al. (2024) CRISPR/Cas9 genome editing of CCR5 combined with C46 HIV-1 fusion inhibitor for cellular resistant to R5 and X4 tropic HIV-1. Sci Rep 14: 10852. https://doi.org/10.1038/s41598-024-61626-x 
|
| [61] |
Spada SJ, Grigg ME, Bouamr F, et al. (2024) TRIM5α: A protean architect of viral recognition and innate immunity. Viruses 16: 997. https://doi.org/10.3390/v16070997 
|
| [62] |
Besnard E, Hakre S, Kampmann M, et al. (2016) The mTOR complex controls hiv latency. Cell Host Microbe 20: 785-797. https://doi.org/10.1016/j.chom.2016.11.001 
|
| [63] |
Henrich TJ, Schreiner C, Cameron C, et al. (2021) Everolimus, an mTORC1/2 inhibitor, in ART-suppressed individuals who received solid organ transplantation: A prospective study. Am J Transplant 21: 1765-1779. https://doi.org/10.1111/ajt.16244 
|
| [64] |
Crater JM, Nixon DF, Furler O'Brien RL (2022) HIV-1 replication and latency are balanced by mTOR-driven cell metabolism. Front Cell Infect Microbiol 12: 1068436. https://doi.org/10.3389/fcimb.2022.1068436 
|
| [65] |
Mu W, Rezek V, Martin H, et al. (2022) Autophagy inducer rapamycin treatment reduces IFN-I-mediated Inflammation and improves anti-HIV-1 T cell response in vivo. JCI Insight 7: e159136. https://doi.org/10.1172/jci.insight.159136 
|
| [66] |
Marković V, Szczepańska A, Berlicki Ł (2024) Antiviral protein-protein interaction inhibitors. J Med Chem 67: 3205-3231. https://doi.org/10.1021/acs.jmedchem.3c01543 
|
| [67] |
Bao Q, Zhou J (2023) Various strategies for developing APOBEC3G protectors to circumvent human immunodeficiency virus type 1. Eur J Med Chem 250: 115188. https://doi.org/10.1016/j.ejmech.2023.115188 
|
| [68] |
McKellar MS (2025) Lenacapavir: a first-in-class capsid inhibitor for HIV treatment and prevention. Curr Opin Infect Dis 38: 208-213. https://doi.org/10.1097/QCO.0000000000001113 
|
| [69] |
Rashid F, Zaongo SD, Iqbal H, et al. (2024) Interactions between HIV proteins and host restriction factors: implications for potential therapeutic intervention in HIV infection. Front Immunol 15: 1390650. https://doi.org/10.3389/fimmu.2024.1390650 
|
| [70] |
Kula-Pacurar A, Rodari A, Darcis G, et al. (2021) Shocking HIV-1 with immunomodulatory latency reversing agents. Semin Immunol 51: 101478. https://doi.org/10.1016/j.smim.2021.101478 
|
| [71] |
Mody A, Sohn AH, Iwuji C, et al. (2024) HIV epidemiology, prevention, treatment, and implementation strategies for public health. Lancet 403: 471-492. https://doi.org/10.1016/S0140-6736(23)01381-8 
|
| [72] |
Pellaers E, Janssens J, Wils L, et al. (2025) BRD4 modulator ZL0580 and LEDGINs additively block and lock HIV-1 transcription. Nat Commun 16: 4226. https://doi.org/10.1038/s41467-025-59398-7 
|
| [73] |
Li C, Mori L, Valente ST (2021) The block-and-lock strategy for human immunodeficiency virus cure: Lessons learned from didehydro-cortistatin A. J Infect Dis 223: 46-53. https://doi.org/10.1093/infdis/jiaa681 
|
| [74] |
Nel C, Frater J (2024) Enhancing broadly neutralising antibody suppression of HIV by immune modulation and vaccination. Front Immunol 15: 1478703. https://doi.org/10.3389/fimmu.2024.1478703 
|
| [75] |
Rossignol E, Alter G, Julg B (2021) Antibodies for human immunodeficiency Virus-1 Cure strategies. J Infect Dis 223: 22-31. https://doi.org/10.1093/infdis/jiaa165 
|
| [76] |
Gieselmann L, DeLaitsch AT, Rohde M, et al. (2025) Profiling of HIV-1 elite neutralizer cohort reveals a CD4bs bnAb for HIV-1 prevention and therapy. Nat Immunol 26: 2016-2029. https://doi.org/10.1038/s41590-025-02286-5 
|
| [77] |
Kopycinski J, Yang H, Hancock G, et al. (2023) Therapeutic vaccination following early antiretroviral therapy elicits highly functional T cell responses against conserved HIV-1 regions. Sci Rep 13: 17155. https://doi.org/10.1038/s41598-023-42888-3 
|
| [78] |
Osborne O, Peyravian N, Nair M, et al. (2020) The paradox of HIV blood-brain barrier penetrance and antiretroviral drug delivery deficiencies. Trends Neurosci 43: 695-708. https://doi.org/10.1016/j.tins.2020.06.007 
|
| [79] |
Li K, Zhang Q (2024) Eliminating the HIV tissue reservoir: current strategies and challenges. Infect Dis (Lond) 56: 165-182. https://doi.org/10.1080/23744235.2023.2298450 
|
| [80] |
Castilha EP, Biondo R, Trugilo KP, et al. (2025) APOBEC3 proteins: From antiviral immunity to oncogenic drivers in HPV-positive cancers. Viruses 17: 436. https://doi.org/10.3390/v17030436 
|
| [81] |
Butler K, Banday AR (2023) APOBEC3-mediated mutagenesis in cancer: causes, clinical significance, and therapeutic potential. J Hematol Oncol 16: 31. https://doi.org/10.1186/s13045-023-01425-5 
|
| [82] |
Said N, Venketaraman V (2025) Neuroinflammation, blood-brain barrier, and HIV reservoirs in the CNS: An in-depth exploration of latency mechanisms and emerging therapeutic strategies. Viruses 17: 572. https://doi.org/10.3390/v17040572 
|